Insulin Action
Insulin is one of the most important hormones in the body. Mutations in the insulin action pathway present substantial increases in longevity in many animals. We use technologies such as global analysis of proteins and their modifications in cells and metabolomics to explore these pathways.
The systems we explore in our analysis of insulin action include reproductive efficiency in flies, glucose transport and GLUT4 translocation to the membrane in adipocytes, protein synthesis in mammalian cells, the circadian rhythm and gene transcription. Our ultimate challenge is to use mathematical skills to integrate these approaches so we can visualise new regulatory features in these complex systems.
Cell Biology of GLUT4 Trafficking
One of the major rate-determining steps for insulin action in muscle and fat (but not liver) cells is glucose transport. Glucose is brought into the cell via GLUT4. Insulin stimulates glucose uptake via the delivery of vesicles containing GLUT4 to the cell surface. A major focus is now on the Rab GTPases that regulate this process since it was discovered that one of the major insulin regulated Akt substrates in muscle and fat cells is a Rab GTPase activating protein, AS160/TBC1D4. We have developed powerful Total Internal Reflection Fluorescence Microscopy (TIRFM) approaches to track the approach, docking and fusion of GLUT4 vesicles at the surface of adipocytes. This has revealed an unexpected heterogeneity in the insulin sensitivity of this process between individual cells. We are currently exploring the basis for this diversity. We have also identified a handful of novel insulin regulated phosphorylation sites in proteins suspected to be involved in GLUT4 trafficking and these also serve as the basis for ongoing projects in the lab.
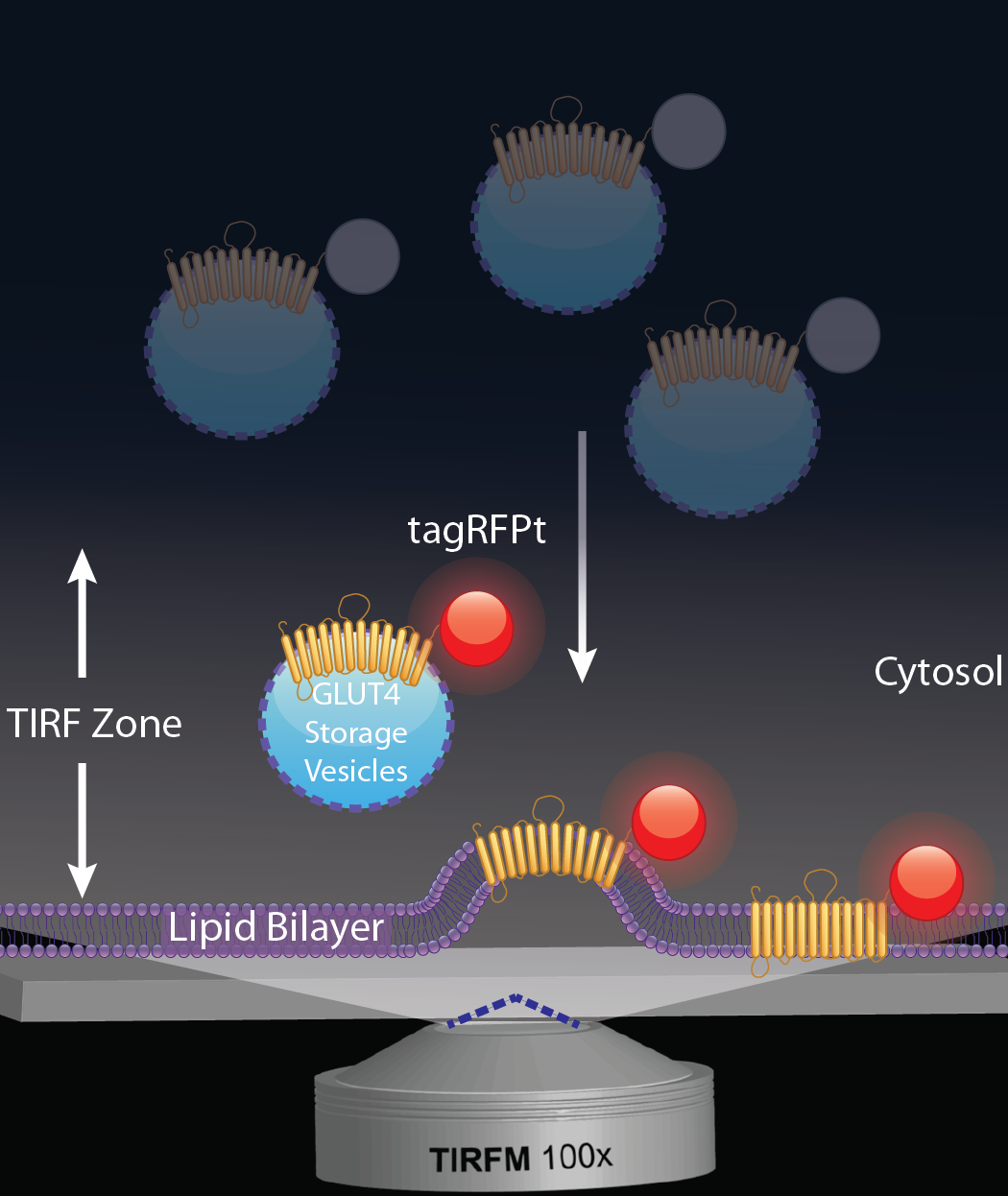
A schematic depicting GLUT4 trafficking with the use of Total Internal Reflection Fluorescence Microscopy (TIRFM)
GLUT4 trafficking in 3T3-L1 Adipocytes is heterogeneous and reproducible
Akt
Akt is a Ser/Thr kinase that sits at a pivotal control point in growth factor signaling. Many growth factor receptors stimulate PI 3’ kinase leading to phosphorylation of Akt at multiple sites commensurate with its activation. Akt phosphorylates numerous substrates integral to many biological processes: glucose transport (AS160/TBC1D4), protein synthesis (mTOR pathway), lipolysis (PDE3b), transcription (Foxo) and glycogen metabolism (GSK3). We are interested in dissecting the sub-architecture of Akt because the quantitative relationship between Akt phosphorylation at its various sites, Akt activity and its ability to phosphorylate its menu of substrates is of great interest and promises to provide novel insights into both metabolic disease and cancer.
Furthermore, Akt is subject to alternate post-translational modifications and the role of these modifications is also of great interest to us as potential cross talk mechanisms with other pathways. Finally, we have used machine learning approaches to discover novel substrates of Akt and we are currently exploring the underlying biochemical outcome of these phosphorylation reactions.
The prioritisation of candidate Akt kinase substrates predicted by machine learning